FDR / \(s\) value on Correlated Null: betahat = zscore, sebetahat = 1
Lei Sun
2017-02-10
Last updated: 2017-12-21
Code version: 6e42447
Introduction
Apply two FDR-controlling procedures, Benjamini–Hochberg 1995 (“BH”) and Benjamini-Yekutieli 2001 (“BY”), and two \(s\) value models, ash and truncash (with the threshold \(T = 1.96\)) to the simulated, correlated null data. The data are obtained from 5 vs 5 GTEx/Liver samples and 10K top expressed genes, and \(1000\) independent simulation trials.
Compare the numbers of false discoveries (by definition, all discoveries should be false) obtained by these four methods, using FDR \(\leq 0.05\) and \(s\)-value \(\leq 0.05\) as cutoffs.
Simulation: \(p\) values for BH and BY procedures; \(\hat\beta = \hat z\), \(\hat s \equiv 1\), for ash and truncash.
\(\hat z\) obtained from the last step of the pipeline.
library(ashr)
source("../code/truncash.R")p = read.table("../output/p_null_liver_777.txt")
t = read.table("../output/t_null_liver_777.txt")
z = read.table("../output/z_null_liver_777.txt")
m = dim(p)[1]
n = dim(p)[2]
fd.bh = fd.by = fd.ash = fd.truncash = c()
for (i in 1:m) {
p_BH = p.adjust(p[i, ], method = "BH")
fd.bh[i] = sum(p_BH <= 0.05)
p_BY = p.adjust(p[i, ], method = "BY")
fd.by[i] = sum(p_BY <= 0.05)
betahat = -as.numeric(z[i, ])
sebetahat = rep(1, n)
fit.ash = ashr::ash(betahat, sebetahat, method = "fdr", mixcompdist = "normal")
fd.ash[i] = sum(ashr::get_svalue(fit.ash) <= 0.05)
fit.truncash = truncash(betahat, sebetahat, t = qnorm(0.975))
fd.truncash[i] = sum(get_svalue(fit.truncash) <= 0.05)
}Result
Simulated under the global null, FWER \(=\) FDR.
Estimated FWER or FDR by BH
fdr.bh = mean(fd.bh >= 1)
fdr.bh[1] 0.046Estimated FWER or FDR by BY
fdr.by = mean(fd.by >= 1)
fdr.by[1] 0.006Estimated FWER or FDR by ash
fdr.ash = mean(fd.ash >= 1)
fdr.ash[1] 0.157Estimated FWER or FDR by truncash
fdr.truncash = mean(fd.truncash >= 1)
fdr.truncash[1] 0.111Happenstance of false discoveries by four approaches
maxcount = max(c(fd.bh, fd.by, fd.ash, fd.truncash))
xlim = c(0, maxcount)
maxfreq = max(c(max(table(fd.bh)), max(table(fd.by)), max(table(fd.ash)), max(table(fd.truncash))))
ylim = c(0, maxfreq)
plot(table(fd.bh), xlab = "Number of False Discoveries / 10K", ylab = "Frequency", main = "Benjamini - Hochberg 1995", xlim = xlim, ylim = ylim)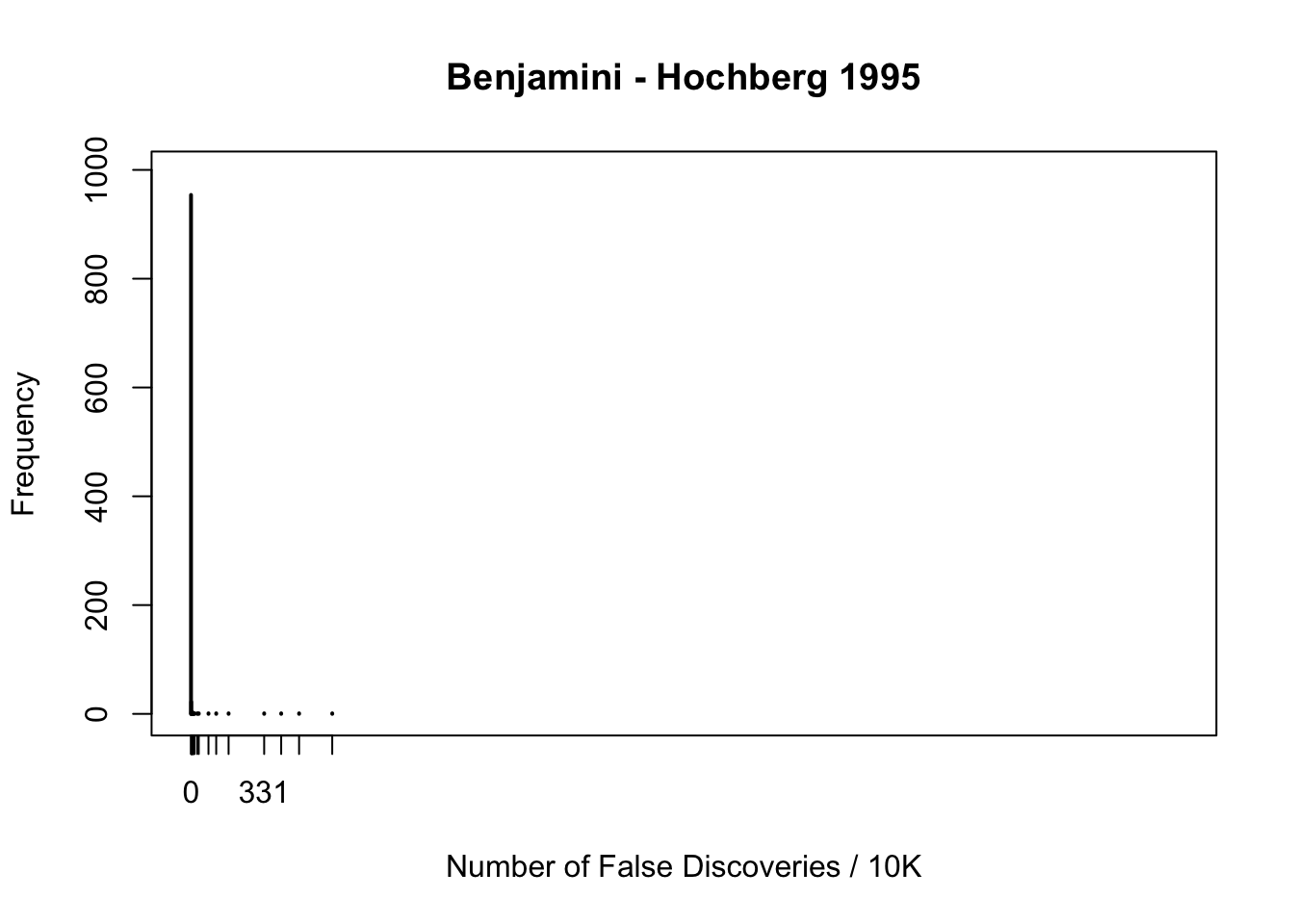
plot(table(fd.by), xlab = "Number of False Discoveries / 10K", ylab = "Frequency", main = "Benjamini - Yekutieli 2001", xlim = xlim, ylim = ylim)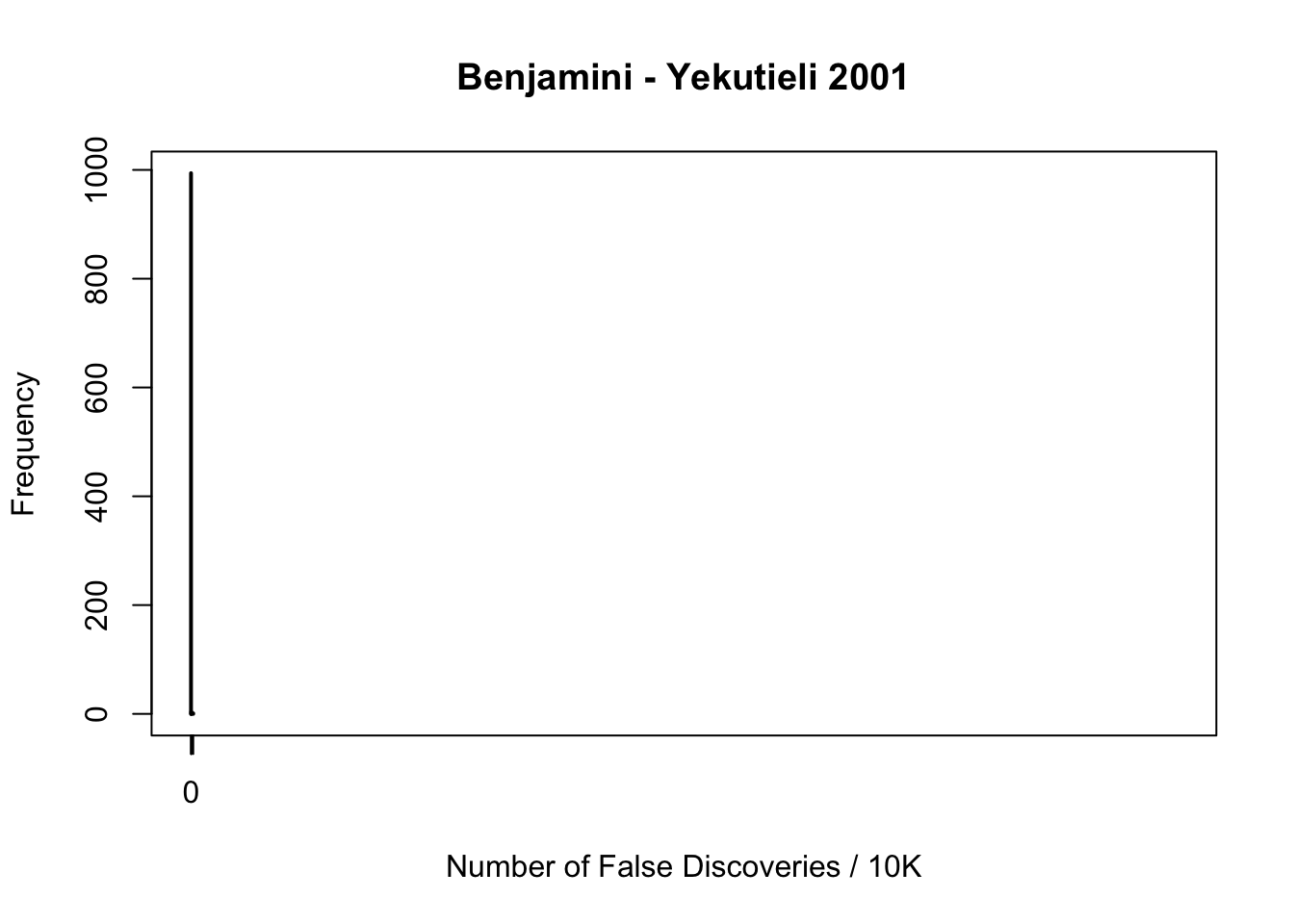
plot(table(fd.ash), xlab = "Number of False Discoveries / 10K", ylab = "Frequency", main = "ash", xlim = xlim, ylim = ylim)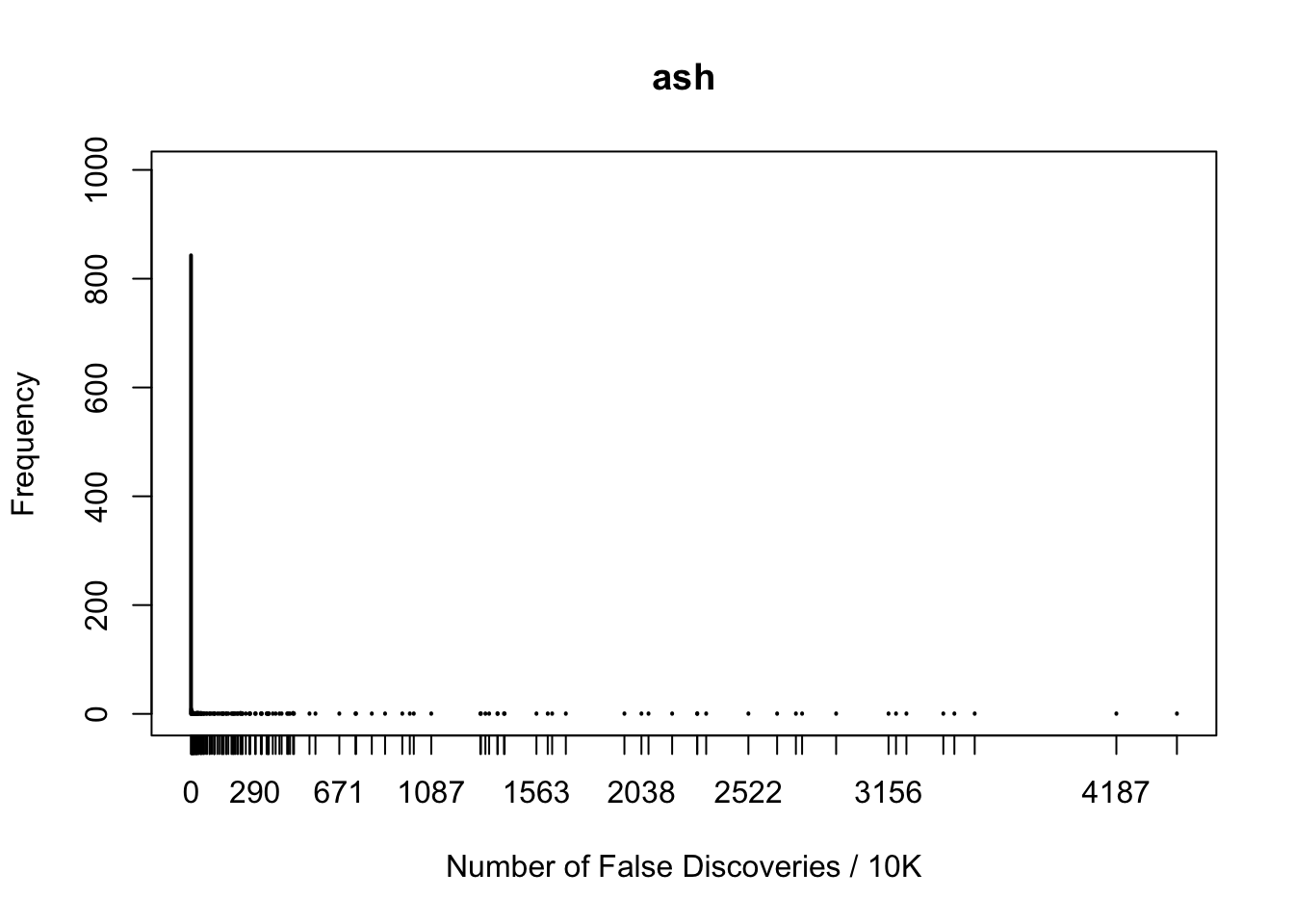
plot(table(fd.truncash), xlab = "Number of False Discoveries / 10K", ylab = "Frequency", main = "truncash", xlim = xlim, ylim = ylim)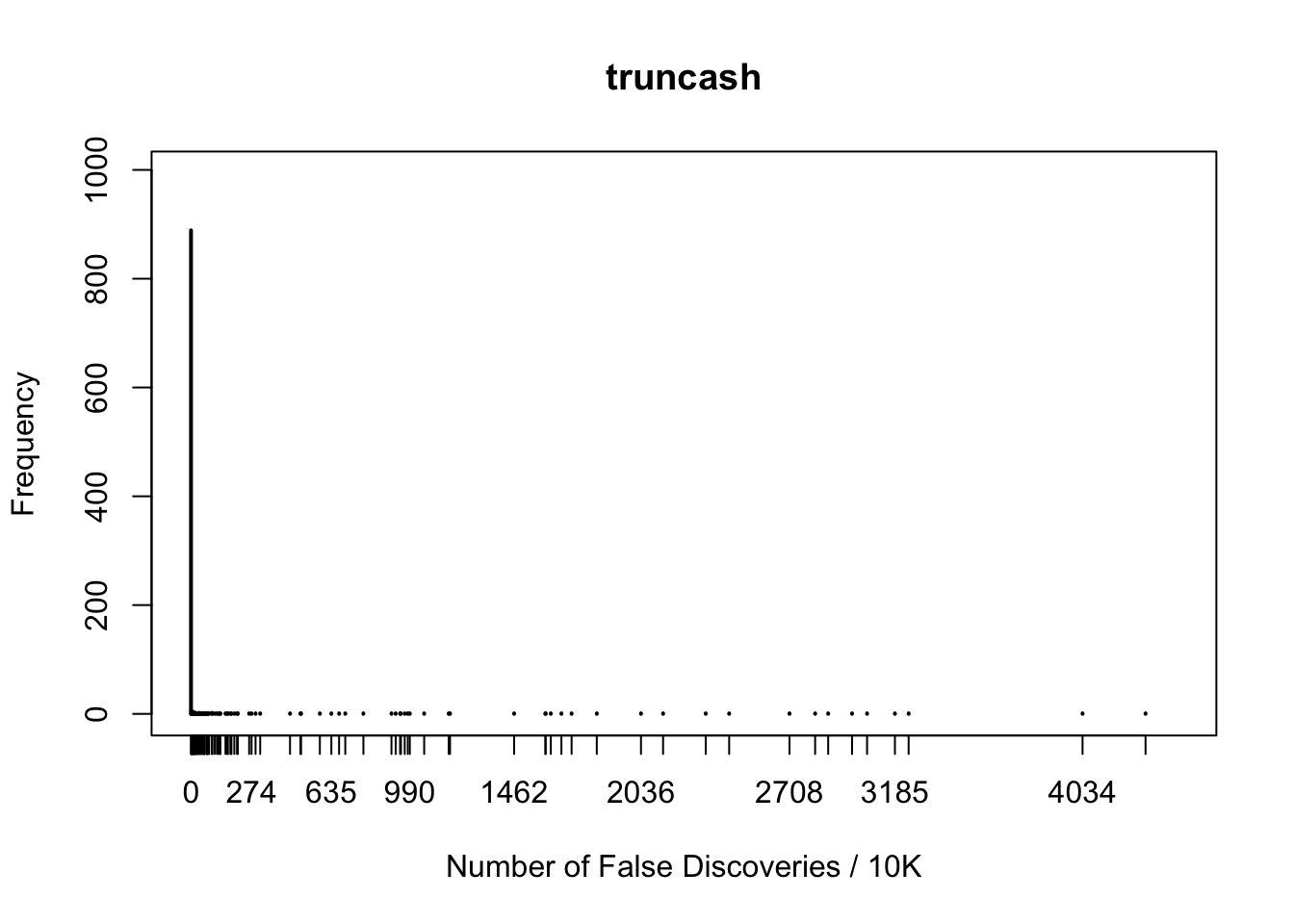
Comparison of the numbers of false discoveries by four approaches
m = length(fd.bh)
fd.ind = (1:m)[!((fd.bh == 0) & (fd.by == 0) & (fd.ash == 0) & (fd.truncash == 0))]
plot(1:length(fd.ind), fd.bh[fd.ind], pch = 4, ylim = xlim, xlab = "Trials with False Discoveries", ylab = "Number of False Discoveries / 10K")
points(1:length(fd.ind), fd.by[fd.ind], pch = 4, col = 2)
points(1:length(fd.ind), fd.ash[fd.ind], pch = 4, col = 3)
points(1:length(fd.ind), fd.truncash[fd.ind], pch = 4, col = 4)
legend("topright", c("BH", "BY", "ash", "truncash"), col = 1:4, pch = 4)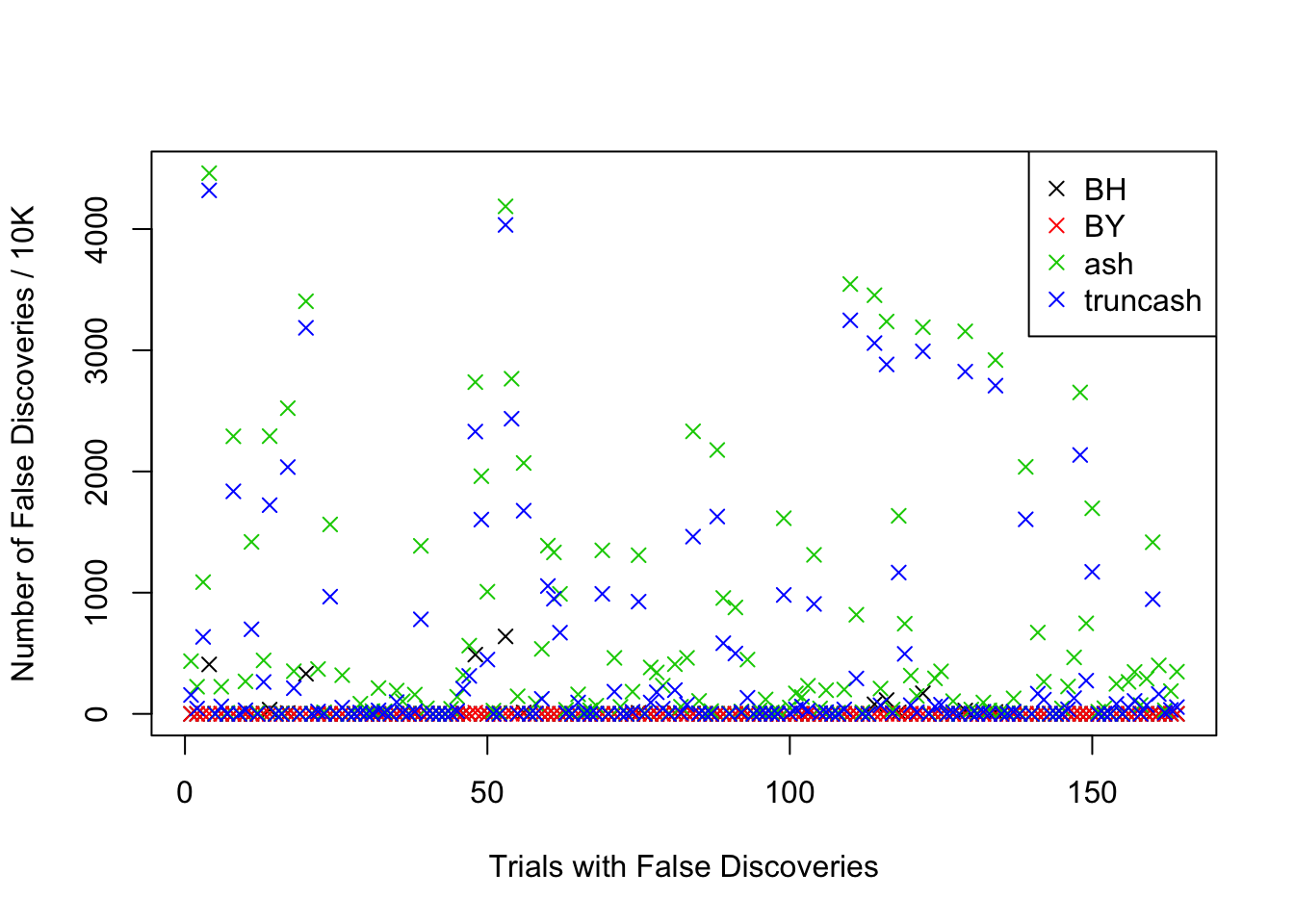
Session Information
sessionInfo()R version 3.4.3 (2017-11-30)
Platform: x86_64-apple-darwin15.6.0 (64-bit)
Running under: macOS High Sierra 10.13.2
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/3.4/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/3.4/Resources/lib/libRlapack.dylib
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] SQUAREM_2017.10-1 ashr_2.2-2
loaded via a namespace (and not attached):
[1] Rcpp_0.12.14 knitr_1.17 magrittr_1.5
[4] workflowr_0.8.0 MASS_7.3-47 doParallel_1.0.11
[7] pscl_1.5.2 lattice_0.20-35 foreach_1.4.4
[10] stringr_1.2.0 tools_3.4.3 parallel_3.4.3
[13] grid_3.4.3 git2r_0.20.0 htmltools_0.3.6
[16] iterators_1.0.9 yaml_2.1.16 rprojroot_1.3-1
[19] digest_0.6.13 Matrix_1.2-12 codetools_0.2-15
[22] evaluate_0.10.1 rmarkdown_1.8 stringi_1.1.6
[25] compiler_3.4.3 backports_1.1.2 truncnorm_1.0-7 This R Markdown site was created with workflowr